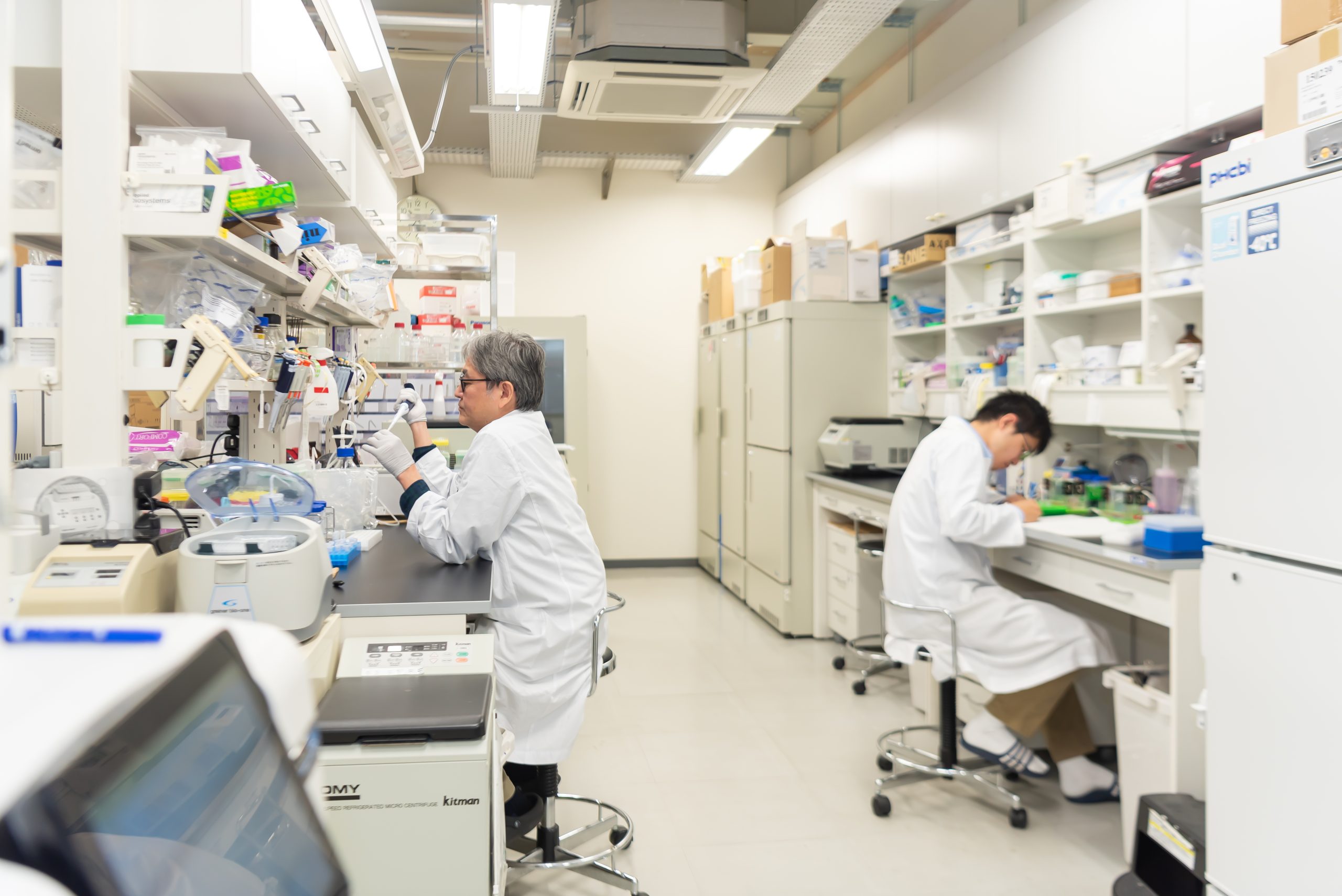
プロジェクト1 – 横山チーム
小児がんや白血病を引き起こす分子メカニズムを解明し新しい薬を生み出す
私たちは、難治性小児がんのメカニズムの解明と分子標的薬の創薬を目指し、特に白血病にフォーカスして研究をしています。白血病は小児がんの主な原因疾患であり、発症には「MYC」というタンパク質が深く関わっています。遺伝子異常の結果、このMYC が異常に産生されることが、がん化を引き起こす主要な原因であることが分かっています。私たちは独自に開発したゲノム解析技術や鶴岡が誇るメタボローム解析を通じて「MYC が引き起こすがん」と「代謝」の関係性を明らかにし、がんの本質に迫りたいと考えています。そして「MYC 関連がん」に対する分子標的薬の開発に取り組み、全ての小児がんに対する解決策を見出したいと思っています。
チームリーダー
横山 明彦Akihiko Yokoyama
1972年愛知県生まれ。
北海道大学理学部化学科卒業。同大学院理学研究科修士課程修了。
東京医科歯科大学医学研究科博士課程修了。
取得学位は、医学博士。スタンフォード大学医学部リサーチアソシエイト、国立がん研究センターユニット長、京都大学医学研究科特定准教授を経て、2016年12月より国立がん研究センター・鶴岡連携研究拠点チームリーダー。
白血病の分子メカニズムに関する研究に従事する。
公益財団法人庄内地域産業振興センター・がんメタボロミクス研究顧問を兼務する。
横山チームチームリーダー
横山 明彦Akihiko Yokoyama

いつか全ての小児がんに対する解決策を
私は、長年MLLという遺伝子に異常を持つ白血病の研究に携わってきました。この白血病は乳児に多く、治りにくいという特徴があり、新たな治療法の開発が切望されています。MLL関連白血病の発症においてMYCというタンパク質が深く関わっています。MYCはよく知られたがん遺伝子であり、遺伝子異常によってMYCファミリータンパク質が異常に産生されることが、様々ながんの原因となっています。神経芽細胞腫など他の難治性小児がんにおいても、MYCが重要な役割を果たしています。私たちはここ鶴岡で「MYC」と「代謝」の関係性を明らかにすることでがんの本質に迫れると考えています。メタボローム(代謝物)解析を通じて、様々な「MYC関連がん」の新たな創薬や診断法の開発に取り組みます。そしてその先にいつか、全ての小児がんに対する解決策を見出したいという夢を持っています。
研究業績
2021
*Yokoyama A. Role of the MOZ/MLL-mediated transcriptional activation system for self-renewal in normal hematopoiesis and leukemogenesis. The FEBS Journal (REVIEW, in press)
Yamamoto K, Goyama S, Asada S, Fujino T, Yonezawa T, Sato N, Takeda R, Tsuchiya A, Fukuyama T, Tanaka Y, Yokoyama A, Toya H, Kon A, Nannya Y, Onoguchi-Mizutani R, Nakagawa S, Hirose T, Ogawa S, Akimitsu N, Kitamura T A histone modifier, ASXL1, interacts with NONO and is involved in paraspeckle formation in hematopoietic cells. Cell Reports 36(8):109576. (2021) doi: 10.1016/j.celrep.2021.109576.
Takahashi S, Kanai A, Okuda H, Miyamoto R, Komata Y, Kawamura T, Matsui H, Inaba T, Takaori-Kondo A, *Yokoyama A. HBO1-MLL interaction promotes AF4/ENL/P-TEFb-mediated leukemogenesis. eLife 10:e65872 (2021) DOI: https://doi.org/10.7554/eLife.65872
Capodanno Y, Chen Y, Schrader J, Tomosugi M, Sumi S, Yokoyama A, Hiraoka N, Ohki R. Cross-talk among MEN1, p53 and Notch regulates the proliferation of pancreatic neuroendocrine tumor cells by modulating INSM1 expression and subcellular localization Neoplasia 23(9):979-992. (2021) doi: 10.1016/j.neo.2021.07.008
*Yokoyama A. Leukemogenesis via aberrant self-renewal by the MLL/AEP-mediated transcriptional activation system. Cancer Science (REVIEW) https://doi.org/10.1111/cas.15054
Miyamoto R, Kanai A, Okuda H, Komata Y, Takahashi S, Matsui H, Inaba T, *Yokoyama A. HOXA9 promotes MYC-mediated leukemogenesis by maintaining gene expression for multiple anti-apoptotic pathways. eLife 10:e64148 (2021) DOI: https://doi.org/10.7554/eLife.64148
Miyamoto R, and Yokoyama A. Protocol for fractionation-assisted native ChIP (fanChIP) to capture protein-protein/DNA interactions on chromatin. STAR Protocols 2, 100404 (2021) doi: https://doi.org/10.1016/j.xpro.2021.100404
2020
Takahashi S, Kanai A, Okuda H, Miyamoto R, Kawamura T, Matsui H, Inaba T, Takaori-Kondo A, *Yokoyama A. HBO1-MLL interaction promotes AF4/ENL/P-TEFb-mediated leukemogenesis. bioRxiv doi: https://doi.org/10.1101/2021.01.08.425834
Miyamoto R, Kanai A, Okuda H, Takahashi S, Matsui H, Inaba T, *Yokoyama A. HOXA9 promotes MYC-mediated leukemogenesis by maintaining gene expression for multiple anti-apoptotic pathways. bioRxiv doi: https://doi.org/10.1101/2020.10.20.347765
Nagamachi A, Kanai A, Nakamura M, Okuda H, Yokoyama A, Shinriki S, Matsui H, Inaba T Multi-organ failure with abnormal receptor metabolism in mice mimicking Samd9/9L syndromes J. Clin. Invest. (2020). https://doi.org/10.1172/JCI140147.
#Miyamoto R, #Okuda H, #Kanai A, Takahashi S, Kawamura T, Matsui H, Kitamura T, Kitabayashi I, Inaba T, *Yokoyama A Activation of CpG-rich promoters mediated by MLL drives MOZ-rearranged leukemia Cell Reports 32:13;108200 (2020) doi.org/10.1016/j.celrep.2020.108200 #co-first author
Takeda R, Asada S, Park SJ, Yokoyama A, Becker H, Kanai A, Visconte V, Hershberger C, Hayashi Y, Yonezawa T, Tamura M, Fukushima T, Tanaka Y, Fukuyama T, Matsumoto A, Yamasaki S, Nakai K, Yamazaki S, Inaba T, Shibata T, Inoue D, Honda H, Goyama S, Maciejewski J, Kitamura T. HHEX promotes myeloid transformation in cooperation with mutant ASXL1 Blood 136 (14): 1670–1684. (2020) DOI:10.1182/blood-2019-126663
Ochi Y, Kon A, Sakata T, Nakagawa MM, Nakazawa N, Kakuta M, Kataoka K, Koseki H, Nakayama M, Morishita D, Tsuruyama T, Saiki R, Yoda A, Okuda R, Yoshizato T, Yoshida K, Shiozawa Y, Nannya Y, Kotani S, Kogure Y, Kakiuchi N, Nishimura T, Makishima H, Malcovati L, Yokoyama A, Takeuchi K, Sugihara E, Sato T, Sanada M, Takaori-Kondo A, Cazzola M, Kengaku M, Miyano S, Shirahige K, Suzuki H, Ogawa S. Combined Cohesin-Runx1 deficiency synergistically perturbs chromatin looping and causes myelodysplastic syndromes. Cancer Discovery 10:836–53 (2020) DOI: 10.1158/2159-8290.CD-19-0982
Takahashi S, *Yokoyama A. The molecular functions of common and atypical MLL fusion protein complexes BBA – Gene Regulatory Mechanisms 1863(7):194548 (2020) [REVIEW] doi: 10.1016/j.bbagrm.2020.194548
Tatsumi G, Kawahara M, Imai T, Nishihata-Asai A, Nishida A, Inatomi O, Yokoyama A, Kakuta Y, Kito K, Andoh A. Thiopurine-mediated impairment of hamtopoietic stem and leukemia cells in Nudt15R138C knock-in mice. Leukemia 34(3):882-894 (2020) DOI ;10.1038/s41375-019-0583-9
Paubelle E, Zylbersztejn F, Maciel TT, Carvalho C, Mupo A, Cheok M, Lieben L, Sujobert P, Decroocq J, Yokoyama A, Asnafi V, Macintyre E, Tamburini J, Bardet V, Castaigne S, Preudhomme C, Dombret H, Carmeliet G, Bouscary D, Ginzburg YZ, de Thé H, Benhamou M, Monteiro RC, Vassiliou GS, Hermine O, Moura IC Vitamin D receptor controls cell stemness in acute myeloid leukemia and in normal bone marrow. Cell Reports. 30(3):739-754. (2020) doi: 10.1016/j.celrep.2019.12.055.
2019
*Yokoyama, A RNA polymerase II-dependent transcription initiated by selectivity factor 1: a central mechanism used by MLL fusion proteins in leukemic transformation (review) Frontiers in Genetics (RNA) 9:722 (2019)
2018
Asada S, Goyama S, Inoue D, Shikata S, Takeda R, Fukushima T, Yonezawa T, Fujino T, Hayashi Y, Kawabata K, Fukuyama T, Tanaka Y, Yokoyama, A Yamazaki S, Kozuka-Hata H, Oyama M, Kojima S, Kawazu M, Mano H, Kitamura T Mutant ASXL1 cooperates with BAP1 to promote myeloid leukaemogenesis Nature Communications 9:2733 (2018)
Inoue D, Fujino T, Sheridan P, Y Zhang Y, Nagase R, Horikawa S, Li Z, Matsui H, Kanai A, Saika M, Yamaguchi R, Kozuka-Hata H, Kawabata K, Yokoyama, A, Goyama S, Inaba T, Imoto S, Miyano S, Xu M, Yang F, Oyama M, and Kitamura T. A novel ASXL1-OGT axis plays roles in H3K4 methylation and tumor suppression in myeloid malignancies Leukemia 32:1327-1337 (2018)
2017
Okuda H, and *Yokoyama, A In vivo leukemogenesis model using retrovirus transduction Bio-protcol 7:23, e2627 (2017)
Okuda H, and *Yokoyama, A Myeloid progenitor transformation assay Bio-protcol 7:23, e2626 (2017)
Chen Y, Anastassiadis K, Kranz A, Stewart AF, Arndt K, Waskow C, Yokoyama A, Jones K, Neff T, Lee Y, Ernst P. MLL2, Not MLL1, Plays a Major Role in Sustaining MLL-Rearranged Acute Myeloid Leukemia. Cancer Cell 31:6, p755-770 (2017)
#Okuda H, #Stanojevic B, Kanai A, Kawamura T, Takahashi S, Matsui H, Takaori-Kondo A, *Yokoyama A. Cooperative gene activation by AF4 and DOT1L drives MLL-rearranged leukemia. J. Clin. Invest. 127(5) 1918-1931#co-first author (2017)
PDF for press release
*Yokoyama, A. Transcriptional activation by MLL fusion proteins in leukemogenesis.(review) Experimental. Hematology 6:21–30 (2017)
2016
Okuda H, Takahashi S, Takaori-Kondo A, *Yokoyama A. TBP loading by AF4 through SL1 is the major rate-limiting step in MLL fusion-dependent transcription. Cell Cycle 15:20, 2712-2722, (2016)
Kadono M, Kanai A, Nagamachi A, Shinriki S, Kanata J, Iwato K, Kyo T, Ochima K, Yokoyama A, Kawamura T, Nagase R, Inoue D, Kitamura T, Inaba T, Ichinohe T, Matsui H. Biological implications of somatic DDX41 p.R525H mutation in acute myeloid leukemia Experimental Hematology 44:745-754 (2016)
2015
Okuda H, Kanai A, Ito S, Matsui H, *Yokoyama A. AF4 utilizes the SL1 components of RNAP1 machinery to initiate MLL fusion- and AEP-dependent transcription Nature Communications 6:8869 (2015)
PDF for press release
*Yokoyama, A. Molecular mechanisms of MLL-associated leukemia.(review) Intl. J. Hematology 101:352-361 (2015).
2014
#Okuda H, #Kawaguchi M, #Kanai A, #Matsui H, Kawamura T, Inaba T, Kitabayashi I, *Yokoyama A. MLL fusion proteins link transcriptional coactivators to previously active CpG-rich promoters Nucleic Acids Research 42:7 4241-4256 (2014) #co-first author
2013
*Yokoyama A, Ficara F, Murphy M, Meisel C, Hatanaka C, Kitabayashi I, *Cleary ML. MLL becomes functional through intra-molecular interaction not by proteolytic processing. PLOS ONE 8(9):e73649 (2013)
2011
*Yokoyama A, Ficara F, Murphy M, Meisel C, Naresh A, Kitabayashi I, *Cleary ML. Proteolytically cleaved MLL subunits are susceptible to distinct degradation pathways. J. Cell Sci. 124(13): 2208-19 (2011)
2010
*Yokoyama A, Lin M, Naresh A, Kitabayashi I, *Cleary ML .A higher-order complex containing AF4- and ENL-family proteins with P-TEFb facilitates oncogenic and physiologic MLL-dependent transcription. Cancer Cell.17(2): 198-212 (2010)
2008
*Yokoyama A and *Cleary ML. Menin critically links MLL proteins with LEDGF on cancer-associated target genes. Cancer Cell. 14:36-46 (2008)
2005
Yokoyama A, Somervaille TCP, Smith KS, Rosenblatt-Rosen O, Meyerson M, Cleary ML .The menin tumor suppressor protein is an essential oncogenic cofactor for MLL-associated leukemogenesis. Cell. 21(123):207-218 (2005)
2004
Yokoyama A, Wang Z, Wysocka J, Sanyal M, Aufiero D, Kitabayashi I, Herr W, Cleary ML. Leukemia proto-oncoprotein MLL forms a SET1-like histone methyltransferase complex with menin to regulate Hox gene expression Mol. Cell. Biol. 24(13):5639-5649 (2004)
2003
Kato K, Yokoyama A, Tohya Y, Akashi H, Nishiyama Y, Kawaguchi Y. Identification of protein kinases responsible for phosphorylation of Epstein–Barr virus nuclear antigen leader protein at serine-35, which regulates its coactivator function J. Gen. Virol. 84;3381-3392 (2003)
2002
Yokoyama A, Kitabayashi I, Ayton PM, Cleary ML, Ohki M. Leukemia proto-oncoprotein MLL is proteolytically processed into two fragments with opposite transcriptional properties. Blood. Nov 15;100(10):3710-8. (2002)
Tanaka M, Yokoyama A, Igarashi M, Matsuda G, Kato K, Kanamori M, Hirai K, Kawaguchi Y, Yamanashi Y. Conserved region CR2 of Epstein-Barr virus nuclear antigen leader protein is a multifunctional domain that mediates self-association as well as nuclear localization and nuclear matrix association. J. Virol. 76(3):1025-32. (2002)
2001
Kitabayashi I, Aikawa Y, Nguyen LA, Yokoyama A, Ohki M. Activation of AML1-mediated transcription by MOZ and inhibition by the MOZ-CBP fusion protein. EMBO J. 20(24):7184-96. (2001)
Yokoyama A, Tanaka M, Matsuda G, Kato K, Kanamori M, Kawasaki H, Hirano H, Kitabayashi I, Ohki M, Hirai K, *Kawaguchi Y. Identification of Major Phosphorylation Sites of Epstein-Barr Virus Nuclear Antigen Leader Protein (EBNA-LP): The function of EBNA-LP to Induce Latent Membrane Protein 1 Cooperatively with EBNA-2 Is Regulated by Phosphorylation. J.Virol. 75(11):5119-5128 (2001)
Kato K, Kawaguchi Y, Tanaka M, Igarashi M, Yokoyama A, Matsuda G, Kanamori M, Nakajima K, Nishimura Y, Shimojima M, Hang TT, Takahashi E, Hirai K Epstein-Barr Virus Encoded-Protein Kinase BGLF4 Mediates Hyper-phosphorylation of Cellular Elongation Factor 1d (EF-1d): EF-1d is Universally Modified by Conserved Protein Kinases of Herpesviruses in Mammalian Cells. J.Gen.Virol. 82(6):1457-63 (2001)
Kawaguchi Y, Tanaka M, Yokoyama A, Matsuda G, Kato K, Kagawa H, Hirai K, and Roizman B. Herpes simplex virus a regulatory protein ICP0 functionally interacts with a cellular transcription factor BMAL1. Proc. Natl. Acad. Sci. USA 98(4) 1877-1882 (2001)
Yokoyama A, Kawaguchi K, Kitabayashi I, Ohki M, Hirai K. The Conserved Domain CR2 of Epstein-Barr Virus Nuclear Antigen Leader Protein is Responsible Not Only for Nuclear Matrix Association But Also for Nuclear Localization. Virology 279, 401-413 (2001)
Kitabayashi I, Aikawa Y, Yokoyama A, Hosoda F, Nagai M, Kakazu N, Abe T, Ohki M. Fusion of MOZ and p300 hitone acetyltransferases in acute monocytic leukemia with a t(8;22)(p11;q13) chromosome translocation. Leukemia 15 89-94 (2001)
2000
Kawaguchi Y, Nakajima K, Igarashi M, Morita T, Tanaka M, Suzuki M, Yokoyama A, Matsuda G, Kato K, Kanamori M, Hirai K. Interaction of epstein-barr virus nuclear antigen leader protein (EBNA-LP) with HS1-associated protein X-1: implication of cytoplasmic function of EBNA-LP. J. Virol. 74(21):10104-11. (2000)
1998
Kitabayashi I, Yokoyama A, Shimizu K, Ohki M. Interaction and functional cooperation of the leukemia-associated factors AML1 and p300 in myeloid cell differentiation. EMBO J. Jun 1;17(11):2994-3004. (1998)
Kitabayashi I, Ida K, Morohoshi F, Yokoyama A, Mitsuhashi N, Shimizu K, Nomura N, Hayashi Y, Ohki M. The AML1-MTG8 leukemic fusion protein forms a complex with a novel member of the MTG8(ETO/CDR) family, MTGR1. Mol. Cell. Biol. 18(2):846-58. (1998)
1997
Yokoyama A, Karasaki H, Urushibara N, Nomoto K, Imai Y, Nakamura K, Mizuno Y, Ogawa K, Kikuchi K. The characteristic gene expressions of MAPK phosphatases 1 and 2 in hepatocarcinogenesis, rat ascites hepatoma cells, and regenerating rat liver. Biochem. Biophys. Res. Commun. 29;239(3):746-51. (1997)
Kaya S, Yokoyama A, Imagawa T, Taniguchi K, Froehlich JP, Albers RW. Cloning of the eel electroplax Na+,K(+)-ATPase alpha subunit. Ann. N Y Acad. Sci. 834:129-31. (1997)
研究概要
(1)がんと代謝
がん細胞は増殖し続けるために必要な栄養を摂取するべく、代謝を変化させる。MYCという遺伝子にコードされるタンパク質は、がん細胞が細胞増殖に適した代謝システムを働かせる上で主要な役割を果たす。我々は、MYCが働く仕組みを明らかにし、MYCが制御するがん特異的な代謝物やその責任酵素を見出すことを目指す。またマウスを使ったがん発症モデルを使って、血中に現れる「がんの最も初期の段階を検出するバイオマーカー」を探索する。これらの研究の成果をもとにがんをできるだけ早期に検出する診断薬や、がん特異的な代謝を阻害する分子標的薬の創製を目指す。
(2)白血病の分子メカニズム
MLLという遺伝子の異常が、悪性度の高い急性白血病を引き起こす。私たちは様々なMLL変異体によって引き起こされる白血病のマウスモデルを構築し、発症のメカニズムを解明することを目指す。MLL遺伝子は染色体転座によって70以上の異なる融合パートナー遺伝子とキメラを形成する。しかし、MLLキメラは70通りの異なるメカニズムで白血病を引き起こすわけではなく、いくつかの基本原理に則った仕組みで、白血病発症を導いている。私たちはMLL関連白血病の基本原理を解明し、その理解に基づいた新たな治療法を発案する。
提供可能な研究技術
■初代培養骨髄前駆細胞を用いたex vivo 培養系
■独自に開発した高感度クロマチン免疫沈降法
研究業績( *corresponding author)
2024
*Yokoyama A, Niida H, *Kutateladze TG, *Côté J. HBO1, a MYSTerious KAT and its links to cancer BBA-Gene Regulatory Mechanisms (REVIEW) (2024)(1867:3:195045) doi.org/10.1016/j.bbagrm.2024.195045
#Gaurav N, #Kanai A, #Lachance C, Cox KL, Liu J, Grzybowski AT, Saksouk N, Klein BJ, Komata Y, Asada S, Ruthenburg AJ, Poirier MG, *Côté J, *Yokoyama A, and *Kutateladze TG. Guiding the HBO1 complex function through the JADE subunit Nature Structural & Molecular Biology
DOI:10.1038/s41594-024-01245-2
https://www.nature.com/articles/s41594-024-01245-2
リリース_白血病を引き起こすタンパク質がゲノムに作用するメカニズムの解明
Becht DC, Kanai A, Halawa M, Biswas S, Shi X, Yokoyama A, and Kutateladze TG. The winged helix domain of MORF binds CpG islands and the TAZ2 domain of p300 iScience
doi.org/10.1016/j.isci.2024.109367
2023
Xiao M, Kondo S, Nomura M, Kato S, Nishimura K, Zhang W, Zhang Y, Akashi T, Viny A, Shigehiro T, Ikawa T, Yamazaki H, Fukumoto M, Tanaka A, Hayashi Y, Koike Y, Aoyama Y, Ito H, Nishikawa H, Kitamura T, Kanai A, Yokoyama A, Fujiwara T, Goyama S, Noguchi H, Lee SC, Toyoda A, Hinohara K, Abdel-Wahab O, Inoue D. BRD9 determines the cell fate of hematopoietic stem cells by regulating chromatin state
Nature Communications (2023) Dec 15;14(1):8372. doi: 10.1038/s41467-023-44081-6.
Chen Y, Yamamoto T, Takahashi Y, Moro T, Tajima T, Sakaguchi Y, Sakata N, Yokoyama A, Hijioka S, Sada A, Tabata Y, Ohki R. Metabolic intervention by low carbohydrate diet suppresses the onset and progression of neuroendocrine tumors Cell Death Disease (2023) Sep 7;14(9):597. doi: 10.1038/s41419-023-06123-1.
#Komata Y, #Kanai A, Maeda T, Inaba T, *Yokoyama A #co-first author, *corresponding author
MOZ/ENL complex is a recruiting factor of leukemic AF10 fusion proteins Nature Communications (2023)14:1979
https://doi.org/10.1038/s41467-023-37712-5
DOI: 10.1038/s41467-023-37712-5
リリース 白血病を引き起こすタンパク質の機能の一端を解明
#Becht DC, #Klein BJ, #Kanai A, #Jang SM, Cox KL, Zhou BR, Phanor SK, Zhang Y1, Chen RW, Ebmeier CC, Lachance C, Galloy M, Fradet-Turcotte A, Bulyk ML, Bai Y, Poirier, MG, *Côté J, *Yokoyama A, and *Kutateladze TG. #co-first author, *co-corresponding author
MORF and MOZ acetyltransferases target unmethylated CpG islands through the winged helix domain Nature Communications (2023)14:697 https://doi.org/10.1038/s41467-023-36368-5
DOI:10.1038/s41467-023-36368-5
白血病細胞が悪性度を維持しながら増殖するメカニズムを解明
2022
*Okuda H, Miyamoto M, Takahashi S, Kawamura T Ichikawa J, Harada I, Tamura T and *Yokoyama A RNA-Binding Proteins of KHDRBS and IGF2BP families control the Oncogenic Activity of MLL-AF4 Nature Communications (2022) 13:6688 DOI:10.1038/s41467-022-34558-1
リリース_悪性度の高い白血病のがん遺伝子発現制御機構を発見
Tanaka A, Nakano T, Nomura M, Yamazaki H, Bewersdorf J, Mulet-Lazaro R, Hogg S, Liu B, Penson A, Yokoyama A, Zang W, Havermans M, Koizumi M, Hayashi Y, Cho H, Kanai A, Lee S, Xiao M, Koike Y, Zhang Y, Fukumoto M, Aoyama Y, Konuma T, Kunimoto H, Inaba T, Nakajima H, Honda H, Kawamoto H, Dell R, Abdel-Wahab O, Inoue D Aberrant EVI1 splicing contributes to EVI1-rearranged leukemia. Blood PMID: 35709354
DOI: 10.1182/blood.2021015325 https://pubmed.ncbi.nlm.nih.gov/35709354/
Koyaguchi T, Niida H, Motegi A, Sakai S, Uchida C, Ohata T, Iijima K, Yokoyama A, Suda T, Kitagawa M. Chromatin-remodeling factor BAZ1/ACF1 targets UV damage sites in an MLL1-dependent manner to facilitate nucleotide excision repair. BBA-Molecular Cell Research (2022)119332
DOI: https://doi.org/10.1016/j.bbamcr.2022.119332
Isobe T, Takagi M, Sato-Otsubo A, Nishimura A, Nagae G, Yamagishi C, Tamura M, Tanaka Y, Asada S, Takeda R, Tsuchiya A, Wang X, Yoshida K, Nannya Y, Ueno H, Akazawa R, Kato I, Mikami T, Watanabe K, Sekiguchi M, Seki M, Kimura S, Hiwatari M, Kato M, Fukuda S, Tatsuno K, Tsutsumi S, Kanai A, Inaba T, Shiozawa Y, Shiraishi Y, Chiba K, Tanaka H, Kotecha RS, Cruickshank MN, Ishikawa F, Morio T, Eguchi M, Deguchi T, Kiyokawa N, Arakawa Y, Koh K, Aoki Y, Ishihara T, Tomizawa D, Miyamura T, Ishii E, Mizutani S, Wilson NK, Göttgens B, Miyano S, Kitamura T, Goyama S, Yokoyama A, Aburatani H, Ogawa S, Takita J Multi-omics analysis defines highly refractory RAS burdened immature subgroup of infant acute lymphoblastic leukemia Nature Communications (2022) 13:4501 DOI:https://doi.org/10.1038/s41467-022-32266-4
Shinriki S, Hirayama M, Nagamachi A, Yokoyama A, Kawamura T, Kanai A, Kawai H, Iwakiri J, Liu R, Maeshiro M, Tungalag S, Tasaki M, Ueda M, Tomizawa K, Kataoka N, Ideue T, Suzuki Y, Asai K, Tani T, Inaba T, and Matsui H DDX41 coordinates RNA splicing and transcriptional elongation to prevent DNA replication stress in hematopoietic cells Leukemia (2022) 36:2605-2620 DOI: https://doi.org/10.1038/s41375-022-01708-9
2021
*Yokoyama A. Role of the MOZ/MLL-mediated transcriptional activation system for self-renewal in normal hematopoiesis and leukemogenesis. The FEBS Journal (REVIEW, in press)
Yamamoto K, Goyama S, Asada S, Fujino T, Yonezawa T, Sato N, Takeda R, Tsuchiya A, Fukuyama T, Tanaka Y, Yokoyama A, Toya H, Kon A, Nannya Y, Onoguchi-Mizutani R, Nakagawa S, Hirose T, Ogawa S, Akimitsu N, Kitamura T A histone modifier, ASXL1, interacts with NONO and is involved in paraspeckle formation in hematopoietic cells. Cell Reports 36(8):109576. (2021) doi: 10.1016/j.celrep.2021.109576.
Takahashi S, Kanai A, Okuda H, Miyamoto R, Komata Y, Kawamura T, Matsui H, Inaba T, Takaori-Kondo A, *Yokoyama A. HBO1-MLL interaction promotes AF4/ENL/P-TEFb-mediated leukemogenesis. eLife 10:e65872 (2021) DOI: https://doi.org/10.7554/eLife.65872
リリース 2021.8.30 _白血病を引き起こすタンパク質間相互作用の発見 final2
Capodanno Y, Chen Y, Schrader J, Tomosugi M, Sumi S, Yokoyama A, Hiraoka N, Ohki R. Cross-talk among MEN1, p53 and Notch regulates the proliferation of pancreatic neuroendocrine tumor cells by modulating INSM1 expression and subcellular localization Neoplasia 23(9):979-992. (2021) doi: 10.1016/j.neo.2021.07.008
*Yokoyama A. Leukemogenesis via aberrant self-renewal by the MLL/AEP-mediated transcriptional activation system. Cancer Science (REVIEW) https://doi.org/10.1111/cas.15054
Miyamoto R, Kanai A, Okuda H, Komata Y, Takahashi S, Matsui H, Inaba T, *Yokoyama A. HOXA9 promotes MYC-mediated leukemogenesis by maintaining gene expression for multiple anti-apoptotic pathways. eLife 10:e64148 (2021) DOI: https://doi.org/10.7554/eLife.64148
リリース_HOXA9がMYCと協調して白血病を引き起こすメカニズム_final
Miyamoto R, and Yokoyama A. Protocol for fractionation-assisted native ChIP (fanChIP) to capture protein-protein/DNA interactions on chromatin. STAR Protocols 2, 100404 (2021) doi: https://doi.org/10.1016/j.xpro.2021.100404
2020
Nagamachi A, Kanai A, Nakamura M, Okuda H, Yokoyama A, Shinriki S, Matsui H, Inaba T Multi-organ failure with abnormal receptor metabolism in mice mimicking Samd9/9L syndromes J. Clin. Invest. (2020). https://doi.org/10.1172/JCI140147.
#Miyamoto R, #Okuda H, #Kanai A, Takahashi S, Kawamura T, Matsui H, Kitamura T, Kitabayashi I, Inaba T, *Yokoyama A Activation of CpG-rich promoters mediated by MLL drives MOZ-rearranged leukemia Cell Reports 32:13;108200 (2020) doi.org/10.1016/j.celrep.2020.108200 #co-first author
リリース_MOZ白血病におけるMLLの役割
Takeda R, Asada S, Park SJ, Yokoyama A, Becker H, Kanai A, Visconte V, Hershberger C, Hayashi Y, Yonezawa T, Tamura M, Fukushima T, Tanaka Y, Fukuyama T, Matsumoto A, Yamasaki S, Nakai K, Yamazaki S, Inaba T, Shibata T, Inoue D, Honda H, Goyama S, Maciejewski J, Kitamura T. HHEX promotes myeloid transformation in cooperation with mutant ASXL1 Blood 136 (14): 1670–1684. (2020) DOI:10.1182/blood-2019-126663
Ochi Y, Kon A, Sakata T, Nakagawa MM, Nakazawa N, Kakuta M, Kataoka K, Koseki H, Nakayama M, Morishita D, Tsuruyama T, Saiki R, Yoda A, Okuda R, Yoshizato T, Yoshida K, Shiozawa Y, Nannya Y, Kotani S, Kogure Y, Kakiuchi N, Nishimura T, Makishima H, Malcovati L, Yokoyama A, Takeuchi K, Sugihara E, Sato T, Sanada M, Takaori-Kondo A, Cazzola M, Kengaku M, Miyano S, Shirahige K, Suzuki H, Ogawa S. Combined Cohesin-Runx1 deficiency synergistically perturbs chromatin looping and causes myelodysplastic syndromes. Cancer Discovery 10:836–53 (2020) DOI: 10.1158/2159-8290.CD-19-0982
Takahashi S, *Yokoyama A. The molecular functions of common and atypical MLL fusion protein complexes BBA – Gene Regulatory Mechanisms 1863(7):194548 (2020) [REVIEW] doi: 10.1016/j.bbagrm.2020.194548
Tatsumi G, Kawahara M, Imai T, Nishihata-Asai A, Nishida A, Inatomi O, Yokoyama A, Kakuta Y, Kito K, Andoh A. Thiopurine-mediated impairment of hamtopoietic stem and leukemia cells in Nudt15R138C knock-in mice. Leukemia 34(3):882-894 (2020) DOI ;10.1038/s41375-019-0583-9
Paubelle E, Zylbersztejn F, Maciel TT, Carvalho C, Mupo A, Cheok M, Lieben L, Sujobert P, Decroocq J, Yokoyama A, Asnafi V, Macintyre E, Tamburini J, Bardet V, Castaigne S, Preudhomme C, Dombret H, Carmeliet G, Bouscary D, Ginzburg YZ, de Thé H, Benhamou M, Monteiro RC, Vassiliou GS, Hermine O, Moura IC Vitamin D receptor controls cell stemness in acute myeloid leukemia and in normal bone marrow. Cell Reports. 30(3):739-754. (2020) doi: 10.1016/j.celrep.2019.12.055.
2019
*Yokoyama A RNA polymerase II-dependent transcription initiated by selectivity factor 1: a central mechanism used by MLL fusion proteins in leukemic transformation (review) Frontiers in Genetics (RNA) 9:722 (2019)
2018
Asada S, Goyama S, Inoue D, Shikata S, Takeda R, Fukushima T, Yonezawa T, Fujino T, Hayashi Y, Kawabata K, Fukuyama T, Tanaka Y, Yokoyama A Yamazaki S, Kozuka-Hata H, Oyama M, Kojima S, Kawazu M, Mano H, Kitamura T Mutant ASXL1 cooperates with BAP1 to promote myeloid leukaemogenesis Nature Communications 9:2733 (2018)
Inoue D, Fujino T, Sheridan P, Y Zhang Y, Nagase R, Horikawa S, Li Z, Matsui H, Kanai A, Saika M, Yamaguchi R, Kozuka-Hata H, Kawabata K, Yokoyama A, Goyama S, Inaba T, Imoto S, Miyano S, Xu M, Yang F, Oyama M, and Kitamura T. A novel ASXL1-OGT axis plays roles in H3K4 methylation and tumor suppression in myeloid malignancies Leukemia 32:1327-1337 (2018)
2017
Okuda H, and *Yokoyama A In vivo leukemogenesis model using retrovirus transduction Bio-protcol 7:23, e2627 (2017)
Okuda H, and *Yokoyama A Myeloid progenitor transformation assay Bio-protcol 7:23, e2626 (2017)
Chen Y, Anastassiadis K, Kranz A, Stewart AF, Arndt K, Waskow C, Yokoyama A, Jones K, Neff T, Lee Y, Ernst P. MLL2, Not MLL1, Plays a Major Role in Sustaining MLL-Rearranged Acute Myeloid Leukemia. Cancer Cell 31:6, p755-770 (2017)
#Okuda H, #Stanojevic B, Kanai A, Kawamura T, Takahashi S, Matsui H, Takaori-Kondo A, *Yokoyama A. Cooperative gene activation by AF4 and DOT1L drives MLL-rearranged leukemia. J. Clin. Invest. 127(5) 1918-1931#co-first author (2017)
悪性度の高い急性白血病のがん化メカニズム
*Yokoyama A. Transcriptional activation by MLL fusion proteins in leukemogenesis.(review) Experimental. Hematology 6:21–30 (2017)
2016
Okuda H, Takahashi S, Takaori-Kondo A, *Yokoyama A. TBP loading by AF4 through SL1 is the major rate-limiting step in MLL fusion-dependent transcription. Cell Cycle 15:20, 2712-2722, (2016)
Kadono M, Kanai A, Nagamachi A, Shinriki S, Kanata J, Iwato K, Kyo T, Ochima K, Yokoyama A, Kawamura T, Nagase R, Inoue D, Kitamura T, Inaba T, Ichinohe T, Matsui H. Biological implications of somatic DDX41 p.R525H mutation in acute myeloid leukemia Experimental Hematology 44:745-754 (2016)
2015
Okuda H, Kanai A, Ito S, Matsui H, *Yokoyama A. AF4 utilizes the SL1 components of RNAP1 machinery to initiate MLL fusion- and AEP-dependent transcription Nature Communications 6:8869 (2015)
PDF for press release
*Yokoyama A. Molecular mechanisms of MLL-associated leukemia.(review) Intl. J. Hematology 101:352-361 (2015).
2014
#Okuda H, #Kawaguchi M, #Kanai A, #Matsui H, Kawamura T, Inaba T, Kitabayashi I, *Yokoyama A. MLL fusion proteins link transcriptional coactivators to previously active CpG-rich promoters Nucleic Acids Research 42:7 4241-4256 (2014) #co-first author
2013
*Yokoyama A, Ficara F, Murphy M, Meisel C, Hatanaka C, Kitabayashi I, *Cleary ML. MLL becomes functional through intra-molecular interaction not by proteolytic processing. PLOS ONE 8(9):e73649 (2013)
2011
*Yokoyama A, Ficara F, Murphy M, Meisel C, Naresh A, Kitabayashi I, *Cleary ML. Proteolytically cleaved MLL subunits are susceptible to distinct degradation pathways. J. Cell Sci. 124(13): 2208-19 (2011)
2010
*Yokoyama A, Lin M, Naresh A, Kitabayashi I, *Cleary ML .A higher-order complex containing AF4- and ENL-family proteins with P-TEFb facilitates oncogenic and physiologic MLL-dependent transcription. Cancer Cell.17(2): 198-212 (2010)
2008
*Yokoyama A and *Cleary ML. Menin critically links MLL proteins with LEDGF on cancer-associated target genes. Cancer Cell. 14:36-46 (2008)
2005
Yokoyama A, Somervaille TCP, Smith KS, Rosenblatt-Rosen O, Meyerson M, Cleary ML .The menin tumor suppressor protein is an essential oncogenic cofactor for MLL-associated leukemogenesis. Cell. 21(123):207-218 (2005)
2004
Yokoyama A, Wang Z, Wysocka J, Sanyal M, Aufiero D, Kitabayashi I, Herr W, Cleary ML. Leukemia proto-oncoprotein MLL forms a SET1-like histone methyltransferase complex with menin to regulate Hox gene expression Mol. Cell. Biol. 24(13):5639-5649 (2004)
2003
Kato K, Yokoyama A, Tohya Y, Akashi H, Nishiyama Y, Kawaguchi Y. Identification of protein kinases responsible for phosphorylation of Epstein–Barr virus nuclear antigen leader protein at serine-35, which regulates its coactivator function J. Gen. Virol. 84;3381-3392 (2003)
2002
Yokoyama A, Kitabayashi I, Ayton PM, Cleary ML, Ohki M. Leukemia proto-oncoprotein MLL is proteolytically processed into two fragments with opposite transcriptional properties. Blood. Nov 15;100(10):3710-8. (2002)
Tanaka M, Yokoyama A, Igarashi M, Matsuda G, Kato K, Kanamori M, Hirai K, Kawaguchi Y, Yamanashi Y. Conserved region CR2 of Epstein-Barr virus nuclear antigen leader protein is a multifunctional domain that mediates self-association as well as nuclear localization and nuclear matrix association. J. Virol. 76(3):1025-32. (2002)
2001
Kitabayashi I, Aikawa Y, Nguyen LA, Yokoyama A, Ohki M. Activation of AML1-mediated transcription by MOZ and inhibition by the MOZ-CBP fusion protein. EMBO J. 20(24):7184-96. (2001)
Yokoyama A, Tanaka M, Matsuda G, Kato K, Kanamori M, Kawasaki H, Hirano H, Kitabayashi I, Ohki M, Hirai K, *Kawaguchi Y. Identification of Major Phosphorylation Sites of Epstein-Barr Virus Nuclear Antigen Leader Protein (EBNA-LP): The function of EBNA-LP to Induce Latent Membrane Protein 1 Cooperatively with EBNA-2 Is Regulated by Phosphorylation. J.Virol. 75(11):5119-5128 (2001)
Kato K, Kawaguchi Y, Tanaka M, Igarashi M, Yokoyama A, Matsuda G, Kanamori M, Nakajima K, Nishimura Y, Shimojima M, Hang TT, Takahashi E, Hirai K Epstein-Barr Virus Encoded-Protein Kinase BGLF4 Mediates Hyper-phosphorylation of Cellular Elongation Factor 1d (EF-1d): EF-1d is Universally Modified by Conserved Protein Kinases of Herpesviruses in Mammalian Cells. J.Gen.Virol. 82(6):1457-63 (2001)
Kawaguchi Y, Tanaka M, Yokoyama A, Matsuda G, Kato K, Kagawa H, Hirai K, and Roizman B. Herpes simplex virus a regulatory protein ICP0 functionally interacts with a cellular transcription factor BMAL1. Proc. Natl. Acad. Sci. USA 98(4) 1877-1882 (2001)
Yokoyama A, Kawaguchi K, Kitabayashi I, Ohki M, Hirai K. The Conserved Domain CR2 of Epstein-Barr Virus Nuclear Antigen Leader Protein is Responsible Not Only for Nuclear Matrix Association But Also for Nuclear Localization. Virology 279, 401-413 (2001)
Kitabayashi I, Aikawa Y, Yokoyama A, Hosoda F, Nagai M, Kakazu N, Abe T, Ohki M. Fusion of MOZ and p300 hitone acetyltransferases in acute monocytic leukemia with a t(8;22)(p11;q13) chromosome translocation. Leukemia 15 89-94 (2001)
2000
Kawaguchi Y, Nakajima K, Igarashi M, Morita T, Tanaka M, Suzuki M, Yokoyama A, Matsuda G, Kato K, Kanamori M, Hirai K. Interaction of epstein-barr virus nuclear antigen leader protein (EBNA-LP) with HS1-associated protein X-1: implication of cytoplasmic function of EBNA-LP. J. Virol. 74(21):10104-11. (2000)
1998
Kitabayashi I, Yokoyama A, Shimizu K, Ohki M. Interaction and functional cooperation of the leukemia-associated factors AML1 and p300 in myeloid cell differentiation. EMBO J. Jun 1;17(11):2994-3004. (1998)
Kitabayashi I, Ida K, Morohoshi F, Yokoyama A, Mitsuhashi N, Shimizu K, Nomura N, Hayashi Y, Ohki M. The AML1-MTG8 leukemic fusion protein forms a complex with a novel member of the MTG8(ETO/CDR) family, MTGR1. Mol. Cell. Biol. 18(2):846-58. (1998)
1997
Yokoyama A, Karasaki H, Urushibara N, Nomoto K, Imai Y, Nakamura K, Mizuno Y, Ogawa K, Kikuchi K. The characteristic gene expressions of MAPK phosphatases 1 and 2 in hepatocarcinogenesis, rat ascites hepatoma cells, and regenerating rat liver. Biochem. Biophys. Res. Commun. 29;239(3):746-51. (1997)
Kaya S, Yokoyama A, Imagawa T, Taniguchi K, Froehlich JP, Albers RW. Cloning of the eel electroplax Na+,K(+)-ATPase alpha subunit. Ann. N Y Acad. Sci. 834:129-31. (1997)